Phosphoinositide 3-kinase
Contributors to Wikimedia projects
 Article Images
Article Images
Phosphatidylinositol 3-kinases (PI 3-kinases or PI3Ks) are a family of enzymes involved in cellular functions such as cell growth, proliferation, differentiation, motility, survival and intracellular trafficking, which in turn are involved in cancer.
| Phosphatidylinositol 3- and 4-kinase | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
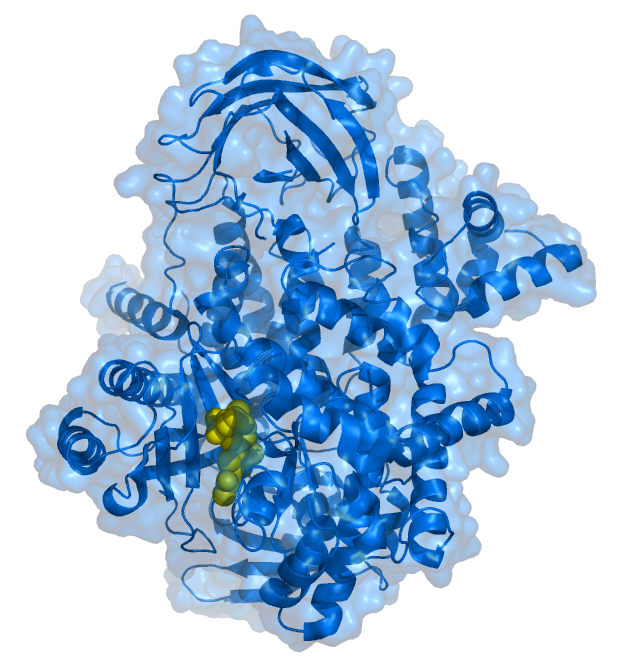
 PI3 Kinase 110 gamma bound to the inhibitor PIK-93 (yellow).[1] | |||||||||||
| Identifiers | |||||||||||
| Symbol | PI3_PI4_kinase | ||||||||||
| Pfam | PF00454 | ||||||||||
| InterPro | IPR000403 | ||||||||||
| SMART | SM00146 | ||||||||||
| PROSITE | PDOC00710 | ||||||||||
| SCOP2 | 3gmm / SCOPe / SUPFAM | ||||||||||
| |||||||||||
PI3Ks are a family of related intracellular signal transducer enzymes capable of phosphorylating the 3 position hydroxyl group of the inositol ring of phosphatidylinositol (PtdIns).[2] They are also known as phosphatidylinositol-3-kinases. The pathway, with oncogene PIK3CA and tumor suppressor PTEN (gene), is implicated in insensitivity of cancer tumors to insulin and IGF1, in calorie restriction.[3][4]
Discovery
The discovery of PI 3-kinases by Lewis Cantley and colleagues began with their identification of a previously unknown phosphoinositide kinase associated with the polyoma middle T protein.[5] They observed unique substrate specificity and chromatographic properties of the products of the lipid kinase, leading to the discovery that this phosphoinositide kinase had the unprecedented ability to phosphorylate phosphoinositides on the 3' position of the inositol ring.[6] Subsequently, Cantley and colleagues demonstrated that in vivo the enzyme prefers PtdIns(4,5)P2 as a substrate, producing the novel phosphoinositide PtdIns(3,4,5)P3.[7]
Classes
PI3Ks interact with the IRS (Insulin receptor substrate) in order to regulate glucose uptake through a series of phosphorylation events.
The phosphoinositol-3-kinase family is divided into three different classes: Class I, Class II, and Class III. The classifications are based on primary structure, regulation, and in vitro lipid substrate specificity.[8]
Class I
Class I PI3Ks are responsible for the production of Phosphatidylinositol 3-phosphate (PI(3)P), Phosphatidylinositol (3,4)-bisphosphate (PI(3,4)P2), and Phosphatidylinositol (3,4,5)-trisphosphate (PI(3,4,5)P3, [<span title="please give a reliable source for this assertion. For Class I, I think it is just PI(3,4,5)P3 which is the product, not PI(3)P or PI(3,4)P2. (January 2010)">citation needed]. The PI3K is activated by G protein-coupled receptors and tyrosine kinase receptors.[8]
Class I PI3K are heterodimeric molecules composed of a regulatory and a catalytic subunit; they are further divided between IA and IB subsets on sequence similarity. Class IA PI3K is composed of a heterodimer between a p110 catalytic subunit and a p85 regulatory subunit.[9] There are five variants of the p85 regulatory subunit, designated p85α, p55α, p50α, p85β, or p55γ. There are also three variants of the p110 catalytic subunit designated p110α, β, or δ catalytic subunit. The first three regulatory subunits are all splice variants of the same gene (Pik3r1), the other two being expressed by other genes (Pik3r2 and Pik3r3, p85β, and p55γ, respectively). The most highly expressed regulatory subunit is p85α; all three catalytic subunits are expressed by separate genes (Pik3ca, Pik3cb, and Pik3cd for p110α, p110β, and p110δ, respectively). The first two p110 isoforms (α and β) are expressed in all cells, but p110δ is expressed primarily in leukocytes, and it has been suggested that it evolved in parallel with the adaptive immune system. The regulatory p101 and catalytic p110γ subunits comprise the type IB PI3K and are encoded by a single gene each.
The majority of the research on PI 3-kinases has focused on the Class I PI 3-kinases. Class I PI 3-kinases are composed of a catalytic subunit known as p110 and a regulatory subunit related to either p85 or p101. The p85 subunits contain SH2 and SH3 domains (Online Mendelian Inheritance in Man (OMIM): 171833). The SH2 domains bind preferentially to phosphorylated tyrosine residues in the amino acid sequence context Y-X-X-M.[10][11]
Classes II and III
Class II and III PI3K are differentiated from the Class I by their structure and function.
Class II comprises three catalytic isoforms (C2α, C2β, and C2γ), but, unlike Classes I and III, no regulatory proteins. Class II catalyse the production of PI(3)P and PI(3,4)P2 from PI; however, little is known about their role in immune cells. C2α and C2β are expressed through the body, however expression of C2γ is limited to hepatocytes. The distinct feature of Class II PI3Ks is the C-terminal C2 domain. This domain lacks critical Asp residues to coordinate binding of Ca2+, which suggests class II PI3Ks bind lipids in a Ca2+-independent manner.
Class III produces only PI(3)P from PI [8] but are more similar to Class I in structure, as they exist as a heterodimers of a catalytic (Vps34) and a regulatory (Vps15/p150) subunits. Class III seems to be primarily involved in the trafficking of proteins and vesicles. There is, however, evidence to show that they are able to contribute to the effectiveness of several process important to immune cells, not least phagocytosis.
Human genes
| group | gene | protein | aliases | EC number |
|---|---|---|---|---|
| class 2 | PIK3C2A | PI3K, class 2, alpha polypeptide | PI3K-C2α | 2.7.1.154 |
| PIK3C2B | PI3K, class 2, beta polypeptide | PI3K-C2β | ||
| PIK3C2G | PI3K, class 2, gamma polypeptide | PI3K-C2γ | ||
| class 3 | PIK3C3 | PI3K, class 3 | Vps34 | 2.7.1.137 |
| catalytic | PIK3CA | PI3K, catalytic, alpha polypeptide | p110-α | 2.7.1.153 |
| PIK3CB | PI3K, catalytic, beta polypeptide | p110-β | ||
| PIK3CG | PI3K, catalytic, gamma polypeptide | p110-γ | ||
| PIK3CD | PI3K, catalytic, delta polypeptide | p110-δ | ||
| regulatory | PIK3R1 | PI3K, regulatory subunit 1 (alpha) | p85-α | N/A |
| PIK3R2 | PI3K, regulatory subunit 2 (beta) | p85-β | ||
| PIK3R3 | PI3K, regulatory subunit 3 (gamma) | p55-γ | ||
| PIK3R4 | PI3K, regulatory subunit 4 | p150 | ||
| PIK3R5 | PI3K, regulatory subunit 5 | p101 | ||
| PIK3R6 | PI3K, regulatory subunit 6 | p87 |
Mechanism
The various 3-phosphorylated phosphoinositides that are produced by PI 3-kinases (PtdIns3P, PtdIns(3,4)P2, PtdIns(3,5)P2, and PtdIns(3,4,5)P3) function in a mechanism by which an assorted group of signalling proteins, containing PX domain, pleckstrin homology domains (PH domains), FYVE domains and other phosphoinositide-binding domains, are recruited to various cellular membranes.
Function
PI 3-kinases have been linked to an extraordinarily diverse group of cellular functions, including cell growth, proliferation, differentiation, motility, survival and intracellular trafficking. Many of these functions relate to the ability of class I PI 3-kinases to activate protein kinase B (PKB, aka Akt) as in the PI3K/AKT/mTOR pathway. The p110δ and p110γ isoforms regulate different aspects of immune responses. PI 3-kinases are also a key component of the insulin signaling pathway. Hence there is great interest in the role of PI 3-kinase signaling in Diabetes mellitus.
Mechanism
The pleckstrin homology domain of AKT binds directly to PtdIns(3,4,5)P3 and PtdIns(3,4)P2, which are produced by activated PI 3-kinase.[12] Since PtdIns(3,4,5)P3 and PtdIns(3,4)P2 are restricted to the plasma membrane, this results in translocation of AKT to the plasma membrane. Likewise, the phosphoinositide-dependent protein kinase 1 (PDK1 or, rarely referred to as PDPK1) also contains a pleckstrin homology domain that binds directly to PtdIns(3,4,5)P3 and PtdIns(3,4)P2, causing it to also translocate to the plasma membrane upon activation of PI 3-kinase. The colocalization of activated PDK1 and AKT allows AKT to become phosphorylated by PDK1 on threonine 308, leading to partial activation of AKT. Full activation of AKT occurs upon phosphorylation of serine 473 by the TORC2 complex of the mTOR protein kinase. (The nomenclature can be confusing. Note that PDK1 also refers to the unrelated enzyme Pyruvate dehydrogenase kinase, isozyme 1. Similarly, TORC2 also refers to the unrelated transcription factor Transducer of Regulated CREB activity 2, which has recently been renamed CREB-regulated transcription coactivator 2 (CRTC2) to reduce the confusion). The "PI3-k/AKT" signaling pathway has been shown to be required for an extremely diverse array of cellular activities - most notably cellular proliferation and survival.
Many other proteins have been identified that are regulated by PtdIns(3,4,5)P3, including Bruton's Tyrosine Kinase (BTK), General Receptor for Phosphoinositides-1 (GRP1), and the O-linked N-acetylglucosamine (O-GlcNAc) transferase.
Cancers
The class IA PI 3-kinase p110α is mutated in many cancers. Many of these mutations cause the kinase to be more active. The PtdIns(3,4,5)P3 phosphatase PTEN that antagonises PI 3-kinase signaling is absent from many tumours. Hence, PI 3-kinase activity contributes significantly to cellular transformation and the development of cancer.
Learning and memory
PI3K has also been implicated in Long-term potentiation (LTP). Whether it is required for the expression or the induction of LTP is still debated. In mouse hippocampal CA1 neurons, PI3K is complexed with AMPA Receptors and compartmentalized at the postsynaptic density of glutamatergic synapses.[13] PI3K is phosphorylated upon NMDA Receptor-dependent CaMKII activity,[14] and it then facilitates the insertion of AMPA-R GluR1 subunits into the plasma membrane. This suggests that PI3K is required for the expression of LTP. Furthermore, PI3K inhibitors abolished the expression of LTP in rat hippocampal CA1, but do not affect its induction.[15] Notably, the dependence of late-phase LTP expression on PI3K seems to decrease over time.[16]
However, another study found that PI3K inhibitors suppressed the induction, but not the expression, of LTP in mouse hippocampal CA1.[17] The PI3K pathway also recruits many other proteins downstream, including mTOR,[18] GSK3β,[19] and PSD-95.[18] The PI3K-mTOR pathway leads to the phosphorylation of p70S6K, a kinase that facilitates translational activity [20] ,[21] further suggesting that PI3K is required for the protein-synthesis phase of LTP induction instead.
PI 3-kinases as protein kinases
Many of the PI 3-kinases appear to have a serine/threonine kinase activity in vitro; however, it is unclear whether this has any role in vivo.
In addition to the class I – class III PI 3-kinases there is a group of more distantly related enzymes that are sometimes referred to as class IV PI 3-kinases. The class IV PI 3-kinases family is composed of ataxia telangiectasia mutated (ATM), ataxia telangiectasia and Rad3 related (ATR), DNA-dependent protein kinase (DNA-PK) and mammalian Target Of Rapamycin (mTOR). These members of the PI 3-kinase superfamily are protein serine/threonine kinases.
Inhibition
All PI 3-kinases are inhibited by the drugs wortmannin and LY294002, although certain members of the class II PI 3-kinase family show decreased sensitivity.
PI 3-kinases inhibitors as therapeutics
As wortmannin and LY294002 are broad inhibitors against PI 3-kinases and a number of unrelated proteins at higher concentrations they are too toxic to be used as therapeutics.[citation needed] A number of pharmaceutical companies have recently been working on PI 3-kinase isoform specific inhibitors including the class I PI 3-kinase, p110δ isoform specific inhibitors, IC486068 and IC87114, ICOS Corporation.[citation needed].GDC-0941 is a highly selective inhibitor of p110α with little activity against mTOR.
References
- ^ PDB: 2chz; Knight ZA, Gonzalez B, Feldman ME, Zunder ER, Goldenberg DD, Williams O, Loewith R, Stokoe D, Balla A, Toth B, Balla T, Weiss WA, Williams RL, Shokat KM (2006). "A pharmacological map of the PI3-K family defines a role for p110alpha in insulin signaling". Cell. 125 (4): 733–47. doi:10.1016/j.cell.2006.03.035. PMC 2946820. PMID 16647110. CS1 maint: multiple names: authors list (link)
- ^ myo-inositol
- ^ Giese N (2009-03-11). "Cell pathway on overdrive prevents cancer response to dietary restriction". PhysOrg.com. Retrieved 2009-04-22.
- ^ Kalaany NY, Sabatini DM (2009). "Tumours with PI3K activation are resistant to dietary restriction". Nature. 458 (7239): 725–31. doi:10.1038/nature07782. PMC 2692085. PMID 19279572.
- ^ Whitman M, Kaplan DR, Schaffhausen B, Cantley L, Roberts TM (1985). "Association of phosphatidylinositol kinase activity with polyoma middle-T competent for transformation". Nature. 315 (6016): 239–42. doi:10.1038/315239a0. PMID 2987699.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Whitman M, Downes CP, Keeler M, Keller T, Cantley L (1988). "Type I phosphatidylinositol kinase makes a novel inositol phospholipid, phosphatidylinositol-3-phosphate". Nature. 332 (6165): 644–6. doi:10.1038/332644a0. PMID 2833705. CS1 maint: multiple names: authors list (link)
- ^ Auger KR, Serunian LA, Soltoff SP, Libby P, Cantley LC (1989). "PDGF-dependent tyrosine phosphorylation stimulates production of novel polyphosphoinositides in intact cells". Cell. 57 (1): 167–75. doi:10.1016/0092-8674(89)90182-7. PMID 2467744. CS1 maint: multiple names: authors list (link)
- ^ a b c Leevers SJ, Vanhaesebroeck B, Waterfield MD (1999). "Signalling through phosphoinositide 3-kinases: the lipids take centre stage". Current Opinion in Cell Biology. 11 (2): 219–25. doi:10.1016/S0955-0674(99)80029-5. PMID 10209156. CS1 maint: multiple names: authors list (link)
- ^ Carpenter CL, Duckworth BC, Auger KR, Cohen B, Schaffhausen BS, Cantley LC (1990). "Purification and characterization of phosphoinositide 3-kinase from rat liver". J. Biol. Chem. 265 (32): 19704–11. PMID 2174051. CS1 maint: multiple names: authors list (link)
- ^ Songyang Z, Shoelson SE, Chaudhuri M; et al. (1993). "SH2 domains recognize specific phosphopeptide sequences". Cell. 72 (5): 767–78. doi:10.1016/0092-8674(93)90404-E. PMID 7680959. ; CS1 maint: multiple names: authors list (link)
- ^ Yoakim M, Hou W, Songyang Z, Liu Y, Cantley L, Schaffhausen B (1994). "Genetic analysis of a phosphatidylinositol 3-kinase SH2 domain reveals determinants of specificity". Mol. Cell. Biol. 14 (9): 5929–38. PMC 359119. PMID 8065326. CS1 maint: multiple names: authors list (link)
- ^ Franke TF, Kaplan DR, Cantley LC, Toker A (1997). "Direct regulation of the Akt proto-oncogene product by phosphatidylinositol-3,4-bisphosphate". Science. 275 (5300): 665–8. doi:10.1126/science.275.5300.665. PMID 9005852. CS1 maint: multiple names: authors list (link)
- ^ Man HY, Wang Q, Lu WY; et al. (2003). "Activation of PI3-kinase is required for AMPA-R insertion during LTP of mEPSCs in cultured hippocampal neurons". Neuron. 38 (4): 611–24. doi:10.1016/S0896-6273(03)00228-9. PMID 12765612. ; CS1 maint: multiple names: authors list (link)
- ^ Joyal JL, Burks DJ, Pons S; et al. (1997). "Calmodulin activates phosphatidylinositol 3-kinase". The Journal of biological chemistry. 272 (45): 28183–6. doi:10.1074/jbc.272.45.28183. PMID 9353264. ; CS1 maint: multiple names: authors list (link) CS1 maint: unflagged free DOI (link)
- ^ Sanna PP, Cammalleri M, Berton F; et al. (2002). "Phosphatidylinositol 3-kinase is required for the expression but not for the induction or the maintenance of long-term potentiation in the hippocampal CA1 region". Journal of Neuroscience. 22 (9): 3359–65. PMID 11978812. ; CS1 maint: multiple names: authors list (link)
- ^ Karpova A, Sanna PP, Behnisch T (2006). "Involvement of multiple phosphatidylinositol 3-kinase-dependent pathways in the persistence of late-phase long term potentiation expression". Neuroscience. 137 (3): 833–41. doi:10.1016/j.neuroscience.2005.10.012. PMID 16326012. CS1 maint: multiple names: authors list (link)
- ^ Opazo P, Watabe AM, Grant SG, O'Dell TJ (2003). "Phosphatidylinositol 3-kinase regulates the induction of long-term potentiation through extracellular signal-related kinase-independent mechanisms". Journal of Neuroscience. 23 (9): 3679–88. PMID 12736339. CS1 maint: multiple names: authors list (link)
- ^ a b Yang PC, Yang CH, Huang CC, Hsu KS (2008). "Phosphatidylinositol 3-kinase activation is required for stress protocol-induced modification of hippocampal synaptic plasticity". The Journal of biological chemistry. 283 (5): 2631–43. doi:10.1074/jbc.M706954200. PMID 18057005. CS1 maint: multiple names: authors list (link) CS1 maint: unflagged free DOI (link)
- ^ Peineau S, Taghibiglou C, Bradley C; et al. (2007). "LTP inhibits LTD in the hippocampus via regulation of GSK3beta". Neuron. 53 (5): 703–17. doi:10.1016/j.neuron.2007.01.029. PMID 17329210. ; CS1 maint: multiple names: authors list (link)
- ^ Toker A, Cantley LC (1997). "Signalling through the lipid products of phosphoinositide-3-OH kinase". Nature. 387 (6634): 673–6. doi:10.1038/42648. PMID 9192891.
- ^ Cammalleri M, Lütjens R, Berton F; et al. (2003). "Time-restricted role for dendritic activation of the mTOR-p70S6K pathway in the induction of late-phase long-term potentiation in the CA1". Proceedings of the National Academy of Sciences of the United States of America. 100 (24): 14368–73. doi:10.1073/pnas.2336098100. PMC 283598. PMID 14623952. ; CS1 maint: multiple names: authors list (link)
Further reading
- Vanhaesebroeck B, Leevers S, Ahmadi K, Timms J, Katso R, Driscoll P, Woscholski R, Parker P, Waterfield M (2001). "Synthesis and function of 3-phosphorylated inositol lipids". Annu Rev Biochem. 70: 535–602. doi:10.1146/annurev.biochem.70.1.535. PMID 11395417.
{{cite journal}}: CS1 maint: multiple names: authors list (link) [1] - Schild C, Wirth M, Reichert M, Schmid RM, Saur D, Schneider G (2009). "PI3K signaling maintains c-myc expression to regulate transcription of E2F1 in pancreatic cancer cells". Mol. Carcinog. 48 (12): 1149–58. doi:10.1002/mc.20569. PMID 19603422. CS1 maint: multiple names: authors list (link)
- Williams; Berndt, A; Miller, S; Hon, WC; Zhang, X; et al. (2009). "Form and flexibility in phosphoinositide 3-kinases". Biochemical Society Transactions. 37 (Pt 4): 615–626. doi:10.1042/BST0370615. PMID 19614567.
External links
- Proteopedia Phosphoinositide_3-Kinases to explore the structure in interactive 3D
- PI-3+Kinase at the U.S. National Library of Medicine Medical Subject Headings (MeSH)